Dans…Orozco {Nucleic Acids Res 44: 4052}

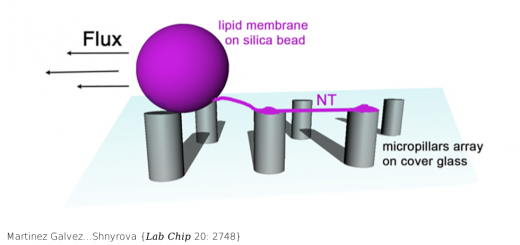
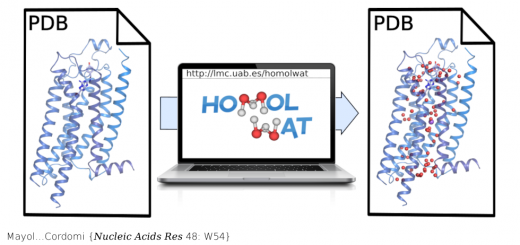
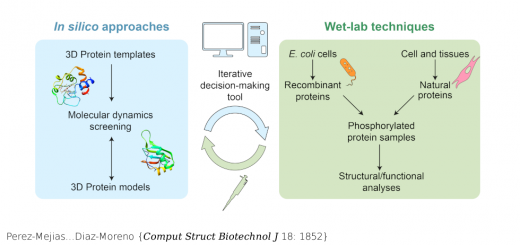
From LEFT to RIGHT: 3D representation of the famous Drew-Dickerson Dodecamer (DDD), the first B-DNA structure solved experimetally (PDB code 1BNA). Major and minor groove views of Potassium distribution along the helix. Cartesian K+ isomolarity surfaces at 1.5 Molar, shown along with the average DNA structure from the MD simulations (represented with a silver sheet). For comparison purposes, the K+ isosurfaces have been overlapped with the Tl+ cations (red spheres) that co-crystallized with the DNA (PDB code 1JGR). Note that Thallium cations are used as a replacement of Potassium in diffraction experiments. The colors of the different isosurfaces represents different state-of-the-art force-field parameterizations available to treat K+ in solution: pink (Jensen & Jorgensen), yellow (Beglov & Roux), green (Joung & Cheatham), blue (Smith & Dang).
Long-timescale dynamics of the Drew-Dickerson dodecamer
Dans PD, Danilāne L, Ivani I, Dršata T, Lankaš F, Hospital A, Walther J, Pujagut RI, Battistini F, Gelpí JL, Lavery R, Orozco M.
Nucleic Acids Res 2016 May; 44: 4052.
We present a systematic study of the long-timescale dynamics of the Drew-Dickerson dodecamer (DDD: d(CGCGAATTGCGC)2) a prototypical B-DNA duplex. Using our newly parameterized PARMBSC1 force field, we describe the conformational landscape of DDD in a variety of ionic environments from minimal salt to 2 M Na(+)Cl(-) or K(+)Cl(-) The sensitivity of the simulations to the use of different solvent and ion models is analyzed in detail using multi-microsecond simulations. Finally, an extended (10 μs) simulation is used to characterize slow and infrequent conformational changes in DDD, leading to the identification of previously uncharacterized conformational states of this duplex which can explain biologically relevant conformational transitions. With a total of more than 43 μs of unrestrained molecular dynamics simulation, this study is the most extensive investigation of the dynamics of the most prototypical DNA duplex.
PubMed: 27084952. Doi: 10.1093/nar/gkw264.